- Home
- Services
- About
- News
- Contact
- Fast and the furious 8 music list
- Chalino sanchez songs las nieves de enero lyrics
- Mashti milano
- Motherload cheats codes
- Epson stylus photo r280 drivers
- How to play dragon age 2 on pc with xbox one controller
- Truecaller app remove person from blocked list
- Doa 5 last round torrent
- How to find mac address on mac os with terminal
- Best structural analysis software
- Free video call recorder for skype echo
- Best free antivirus 2018 for android
- Cydia impactor cpp 81
- Harmor vst presets reddit
- Backuptrans crack keygen
- What is best audio editor for mac
- Men in black 3 full movie for free 213
- Vshare download error
- Urc remote config software windows
- Minecraft server mac 1-8
- Ftp client mac os sierra
- Sandisk flash drive format tool download
- 3d sexvilla 2 ever lust
- Adele 25 cd play list
- How to add footnote reference in word
- Evangelion shinji ikari raising
- Does adobe photoshop elements 11 work with windows 8
- Nhl 2004 rebuilt 2015 crashes after quitting game
- Ps3 game genie codes reide t evil hd
- Microsoft word vertical alignment table
- Easy free home budget spreadsheet
- Best new games 2016 ps4
- Get word for mac
- Play gang beasts no download
- Why does aquamacs use so much cpu
- Blackmagic disk speed test windows version
- Best external webcam for mac
- Viva pinata trouble in paradise wiki jelly
- Cube world free play no download
- License authorization wizard to contact spss inc
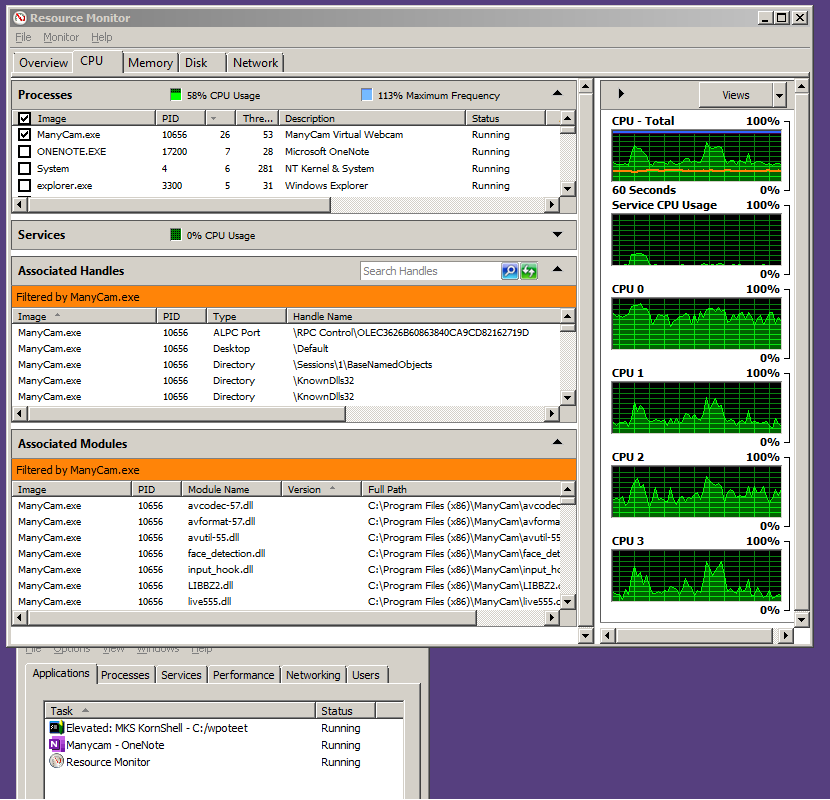
NARR has its share of problems in my region of interest (too hot in the summer, too cold in the winter, winds are often iffy onshore), but nothing so drastic as to make my runs too terrible.īut at 1 km, I have some serious weirdness afoot that didn't appear in the 10 km version. Both runs are surface forced with NARR data (temp, pressure, humidity, precip, winds, radiation, clouds) as put through the ROMS BULK_FLUXES formulation. My cppdefs are otherwise very similar to the 10 km run (which was forced with SODA boundary conditions) U3H/C4V TS advection, standard UV_ADV (which I believe currently defaults to 3rd order horizontal upwinding, 4th order centered in the vertical), LMD vertical mixing, Chapman boundary conditions for zeta and Flather for the mixing TKE. I generated a reasonable 10 km horizontal resolution run for the US west coast that I'm using to provide (mostly) Rad/Nud boundary conditions for my vastly zoomed in 1 km horizontal resolution run, interpolated with the very nice MATLAB scripts provided with the ROMS package these days. I use ROMS/TOMS version 3.6, downloaded from the SVN repository. Since I'm pretty much self-taught from the forums, this does not come as a huge surprise to me, but it's definitely getting to be time to look for help. Or maybe someone else will.I feel like I must be doing something fundamentally wrong in my application of ROMS to the northern California current at very high resolution.
I will make this a separate post once I get time to write out the details. So the URL, using WebEnv and QueryKey, from the previous query looks in my example looks like: For the second query we use the esummary utility to retrieve the actual records. XmlValue(xmlChildren(xmlRoot(parsed.data))$WebEnv)Ģ. XmlValue(xmlChildren(xmlRoot(parsed.data))$QueryKey) # Parse the result for WebEnv and QueryKey: For the first query we use the esearch utility to retrieve two important variables, WebEnv and QueryKey.įor the search term "h1n1" the URL needs to look like: To map keywords by publication date (like in the NYT example) we need to submit two different queries:ġ. Results are in xml, so we'd need the XML library. So all we'd need to do is concatenate the appropriate URL string then parse the results in R. Ricardo Pietrobon suggested to try the above on Pubmed. Plot (freqs, type="l", xaxt="n", main=paste("Search term(s):",q), ylab="# of articles", xlab="date") # can't seem to be able to compare Posix objects with %in%, so coerce them to character for this:įreqs <- ifelse(as.character(dat.all) %in% as.character(strptime(cts$dat, format="%Y-%m-%d")), cts$Freq, 0) # (take out PSD at the end to make it comparable)ĭat.all <- strptime(dat.all, format="%Y-%m-%d") # assign 0 where there is no count, otherwise take count # compare dates from counts dataframe with the whole data range # aggregate counts for dates and coerce into a data frame Uri <- paste0("", q, "&page=", i, "&fl=pub_date&api-key=", api)ĭat <- append(dat, unlist(res$response$docs)) # convert the dates to a vector and appendĭat.conv <- strptime(dat, format="%Y-%m-%d") # need to convert dat into POSIX formatĭaterange <- c(min(dat.conv), max(dat.conv))ĭat.all <- seq(daterange, daterange, by="day") # all possible days Records <- 500 #how many results do we want? (Note API limitations) Q <- "ebola+outbreak" # Query string, use + instead of space There is also an R package by Scott Chamberlain which works with the new API2.Īpi <- "XXXXXXXX" #<<<<<<<<<<<<<= API key goes here Many thanks to Clara Suong for suggesting the changes. This is a revised version which works with the new API2 released in July 2014.
WHY DOES AQUAMACS USE SO MUCH CPU REGISTRATION
To run the code below a (free) registration for an API key is required. It retrieves the publishing dates of articles that contain a query string and plots the number of articles over time, like this: NYT Articles containing "Ebola Outbreak" This is a quick and dirty attempt to make use of the NYT Article Search API from within R.
- Home
- Services
- About
- News
- Contact
- Fast and the furious 8 music list
- Chalino sanchez songs las nieves de enero lyrics
- Mashti milano
- Motherload cheats codes
- Epson stylus photo r280 drivers
- How to play dragon age 2 on pc with xbox one controller
- Truecaller app remove person from blocked list
- Doa 5 last round torrent
- How to find mac address on mac os with terminal
- Best structural analysis software
- Free video call recorder for skype echo
- Best free antivirus 2018 for android
- Cydia impactor cpp 81
- Harmor vst presets reddit
- Backuptrans crack keygen
- What is best audio editor for mac
- Men in black 3 full movie for free 213
- Vshare download error
- Urc remote config software windows
- Minecraft server mac 1-8
- Ftp client mac os sierra
- Sandisk flash drive format tool download
- 3d sexvilla 2 ever lust
- Adele 25 cd play list
- How to add footnote reference in word
- Evangelion shinji ikari raising
- Does adobe photoshop elements 11 work with windows 8
- Nhl 2004 rebuilt 2015 crashes after quitting game
- Ps3 game genie codes reide t evil hd
- Microsoft word vertical alignment table
- Easy free home budget spreadsheet
- Best new games 2016 ps4
- Get word for mac
- Play gang beasts no download
- Why does aquamacs use so much cpu
- Blackmagic disk speed test windows version
- Best external webcam for mac
- Viva pinata trouble in paradise wiki jelly
- Cube world free play no download
- License authorization wizard to contact spss inc
